¶ Introduction
The plugin for connecting ChimeraX to 3decision services, allows users to search for structures, browse projects, and visualize molecular data directly within ChimeraX. More specifically, this version includes:
- Structure Search: Search the 3decision database for molecular structures with real-time job polling
- Project Browser: Browse and load structures from 3decision projects with transformation matrices
- Authentication Management: Secure storage of API credentials with automatic token refresh
- ChimeraX Integration: Native integration with ChimeraX's tool system and menu structure
¶ Installation
¶ Prerequisites
- ChimeraX 1.10.1 or later - Download from ChimeraX official website - might work with older versions as well.
- Active 3decision API credentials (API key and server URL)
¶ Recommended Installation Method
Open Chimera. In the command bar in the bottom of the window type:
toolshed install threedecision
The plugin will then be available in the menu Tools → Structure Analysis → Discngine 3decision
¶ Set up
When opening the plugin for the first time you won't be able to run a search immediately. You first need to enter credentials in order to connect to your 3decision instance.
Click on the cog icon on the top right of the dialog to open the settings menu.
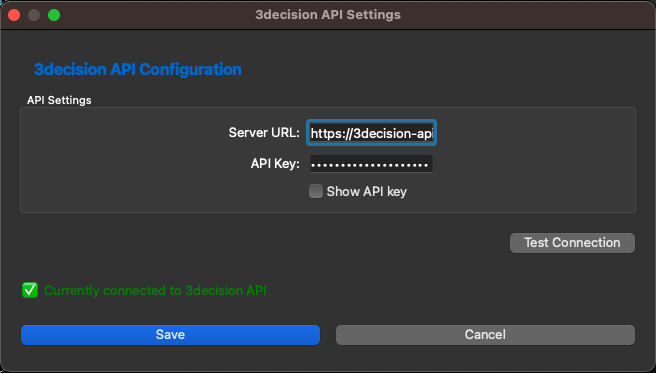
There fill in the URL of the API of 3decision (careful, this URL is different from the usual 3decision URL you are used to). You can find how to generate the API Key you need to insert into this dialog and the URL from this documentation https://3decision-help.discngine.cloud/en/API/Access. NB: The API Key you enter here should be ONLY used by you. Every user needs to set up his own key.
Once you filled both items you can click on Test Connectionwhich will check the authentication vs 3decision.
Don't hesitate to reach out to 3decision support if you have trouble setting this up.
¶ Search
Once you are connected to 3decision, you can start searching. In the search field all text that is supported by 3decision is supported here as well, so you can use excerpts from titles, pdb codes, protein or gene names and identifiers that you used to label your structures or compounds and smiles strings.
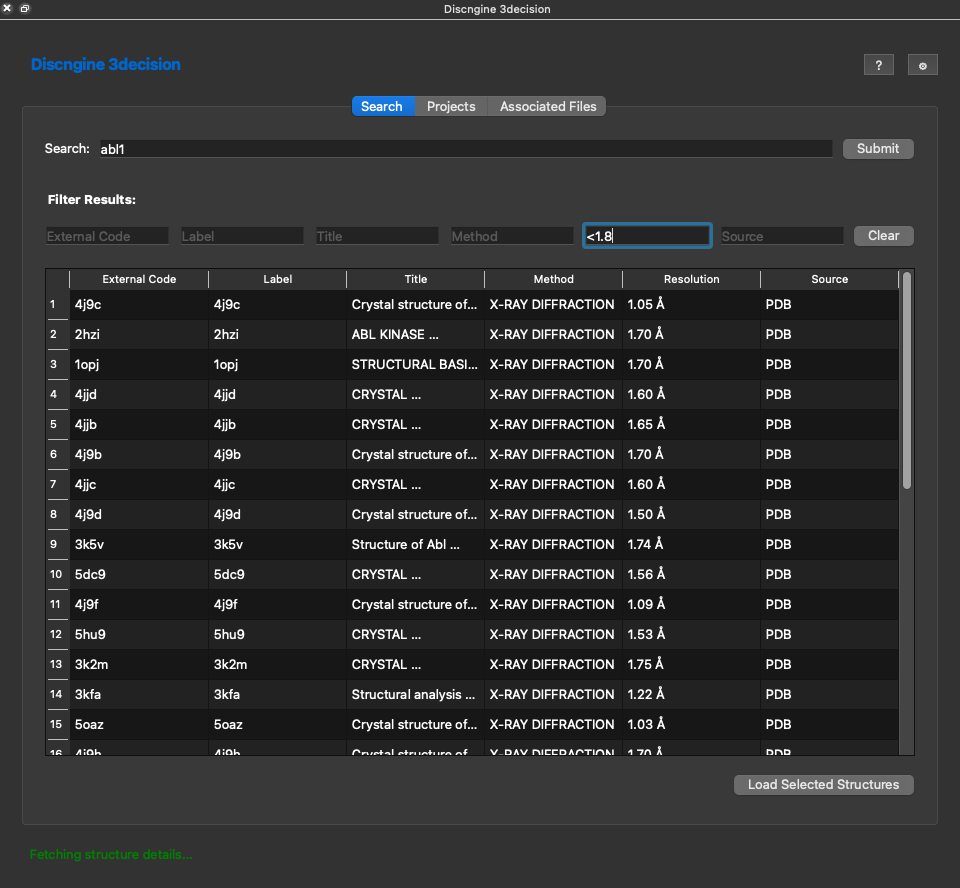
Here for instance I searched for abl1. Hundreds of structures show up in the listing. You can narrow down your search with the filters above each column. For instance if I want to have structures with a resolution better than 1.8 I can type <1.8. The other filters are text filters that are applied while you type to the entries of the table. Note that each column can be sorted as well. Simply click on it for ascending sort and click again for descending sorting.
Once you found the structure(s) you are interested in you can select it in the table and click on Load Selected Structures
This loads the structure into ChimeraX. 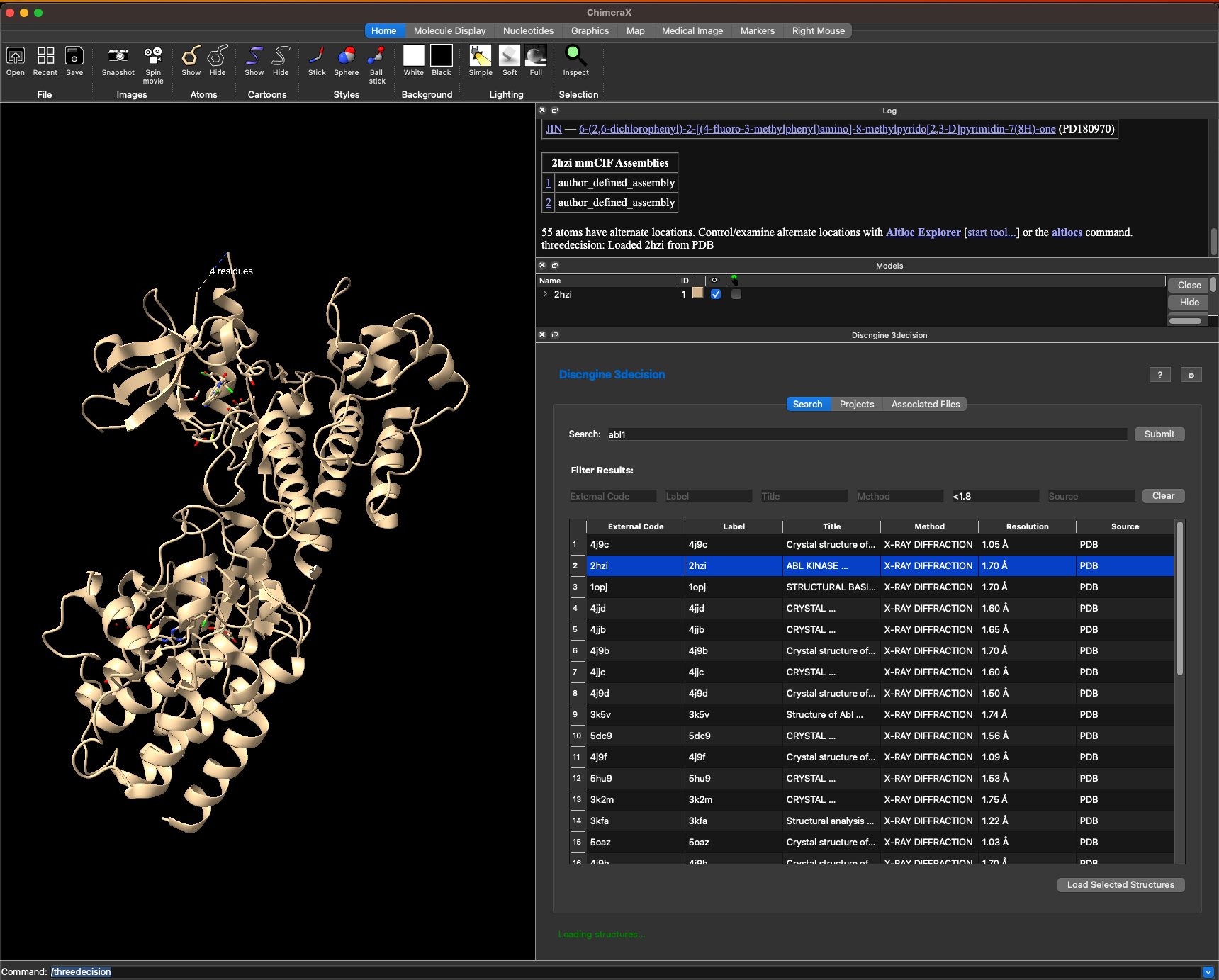
¶ Projects
When clicking on the projects tab you'll see a listing of all projects you have access to, how many structures are in the project and who owns the project.
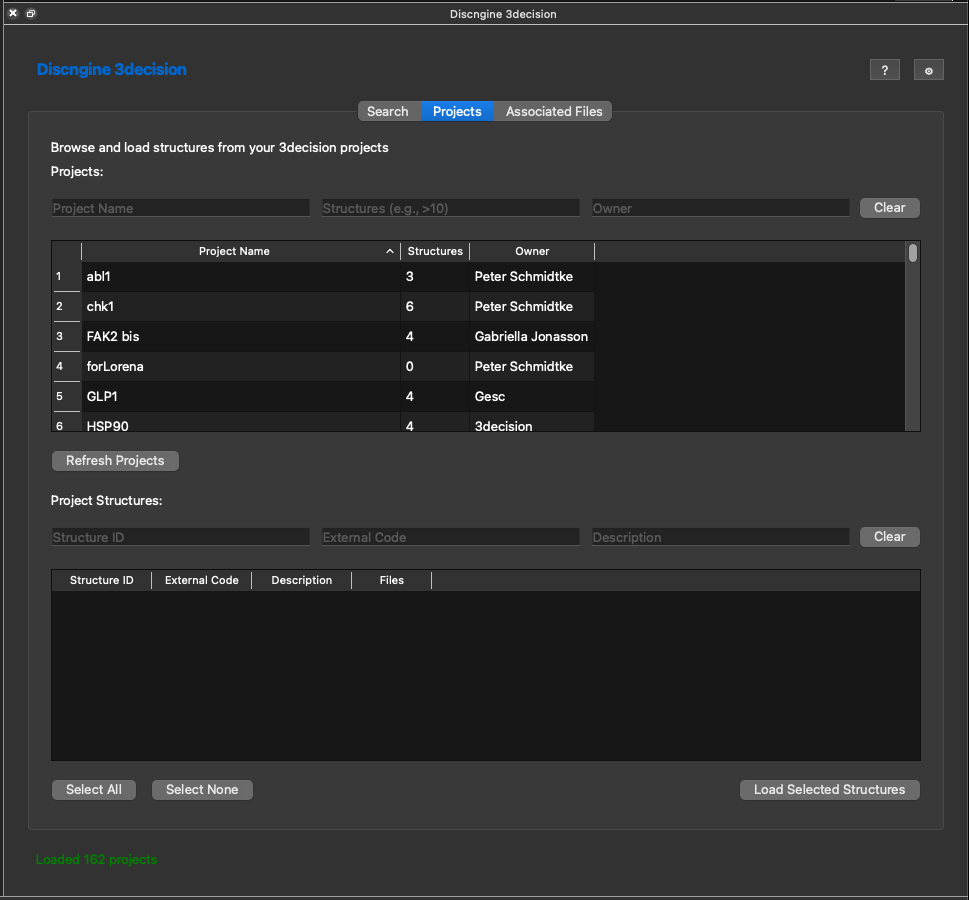
The way as in the search results list, you can filter and sort the project list. When clicking on a particular project you'll see all structures available in the project.
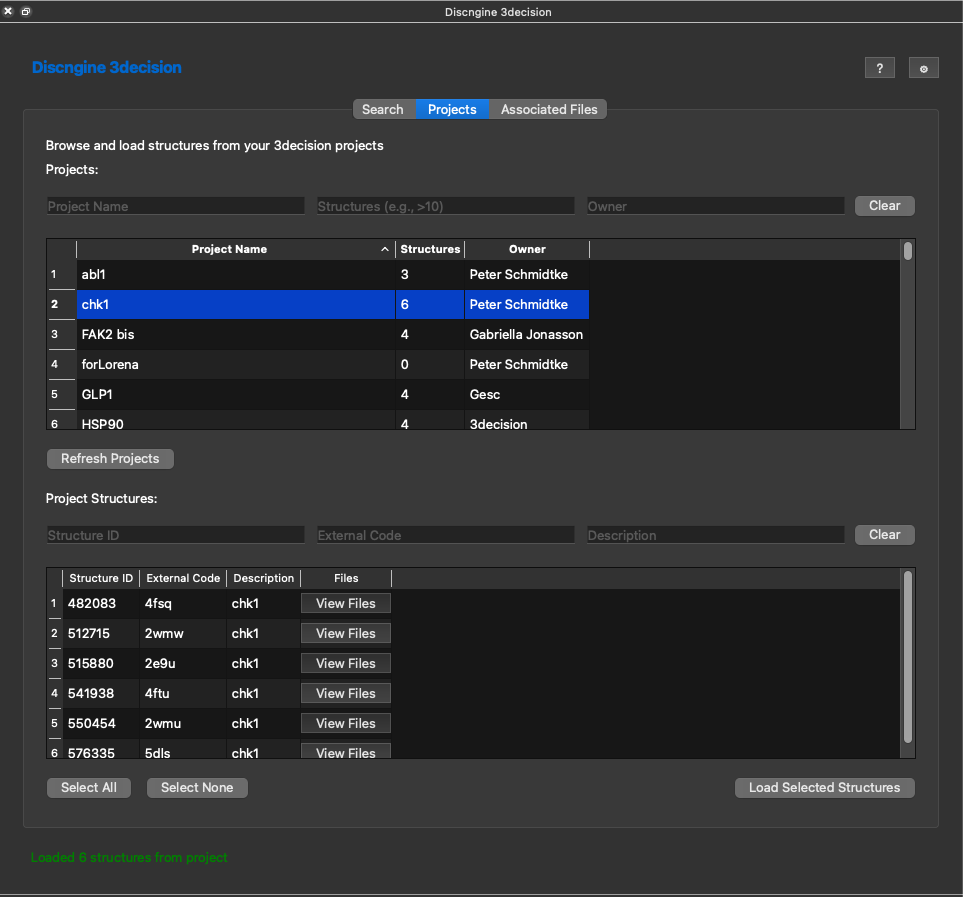
Again in the bottom you can sort and filter. You can select one or several structures here and load these into ChimeraX now.
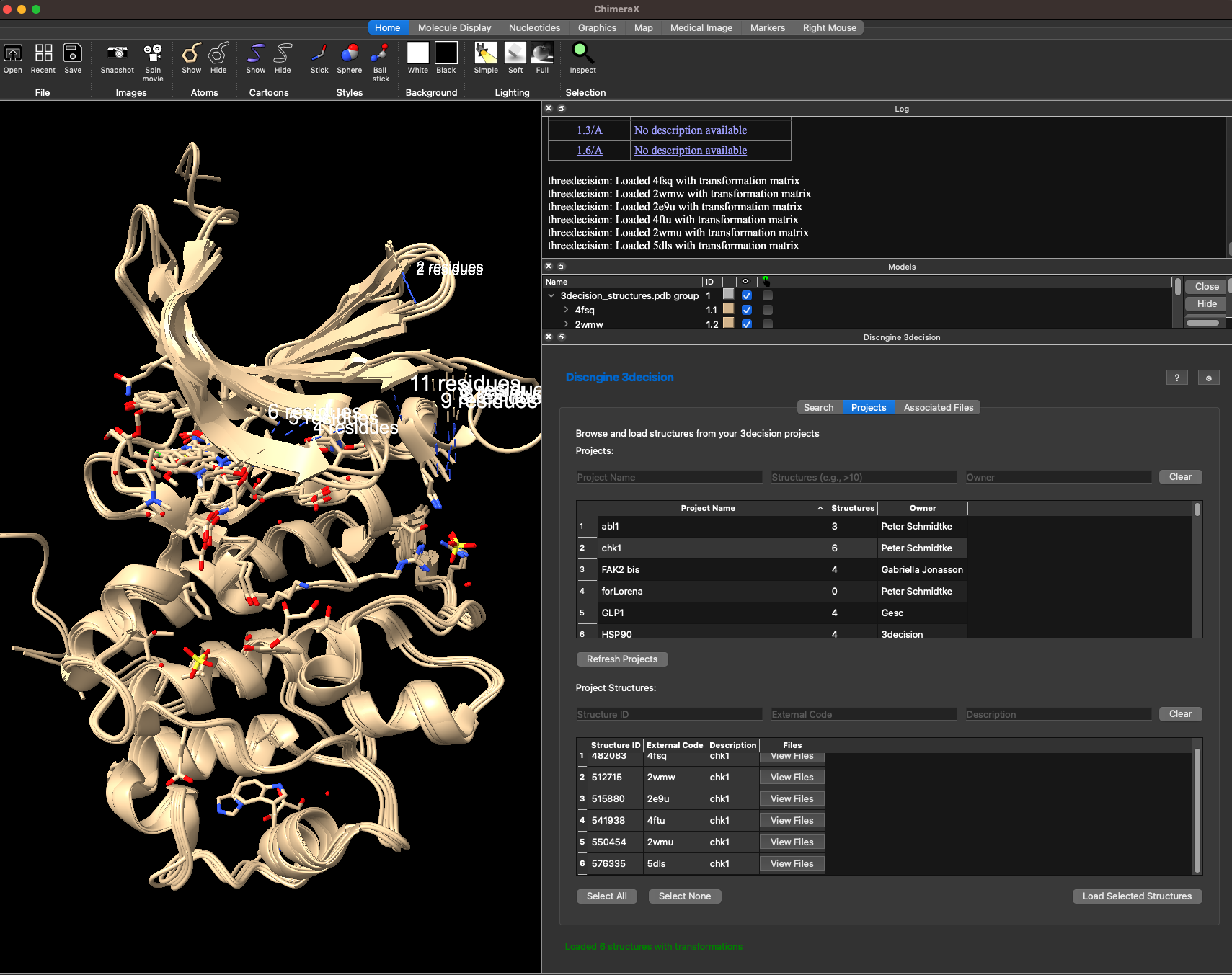
Here I loaded all six structures from my project. You might notice that all structures are nicely superposed on the ATP binding site here. This is because this 3decision project has a common reference structure defined and all structures in the project are automatically aligned when loaded. If you want to know more on how to set this up for your own projects, then you can read more about that here: https://3decision-help.discngine.cloud/en/FeaturesDescription/Projects
¶ Associated Files
When opening structures from projects you might have seen the additional button in the table named View Files. When clicking this button you'll see a listing of all files that are attached to a structure in 3decision.
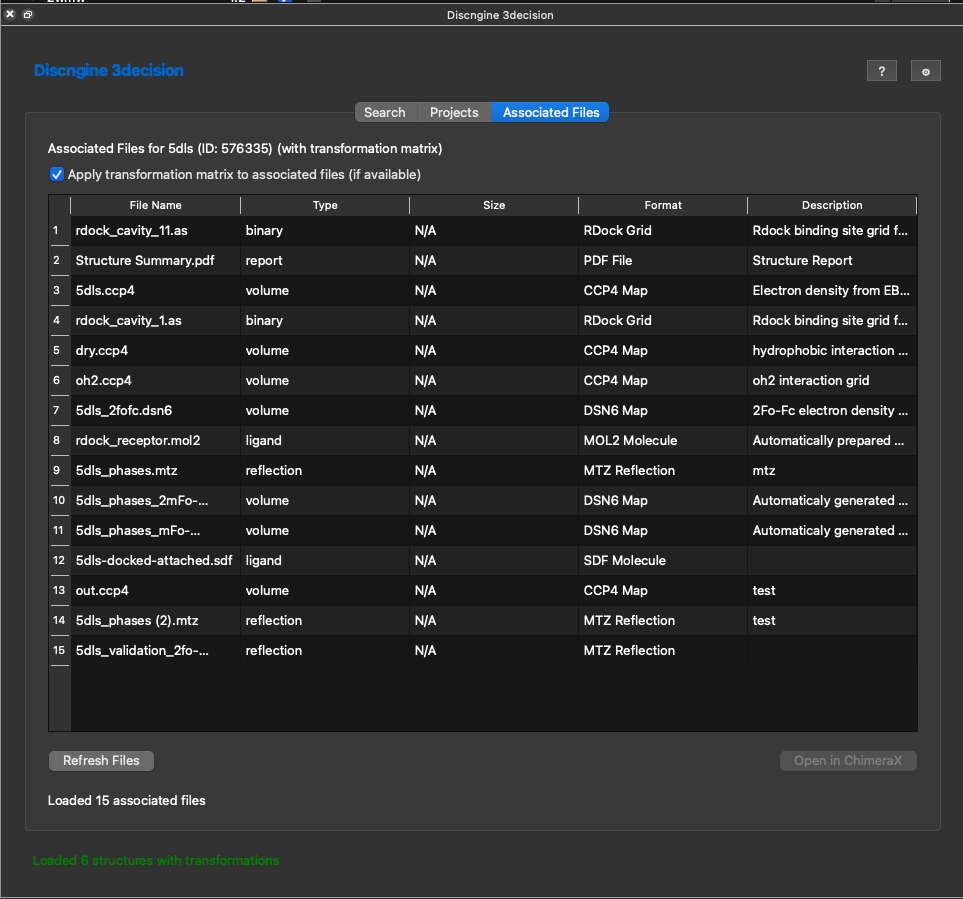
Here are all associated files fro 5dls for instance. You can open the compatible file formats with ChimeraX by selecting one and cliking on Open in ChimeraX.
If you defined a common reference for your project, this associated file will also be transformed to the common reference frame.
Here for instance I opened the ccp4 map of 5dls to the viewer. 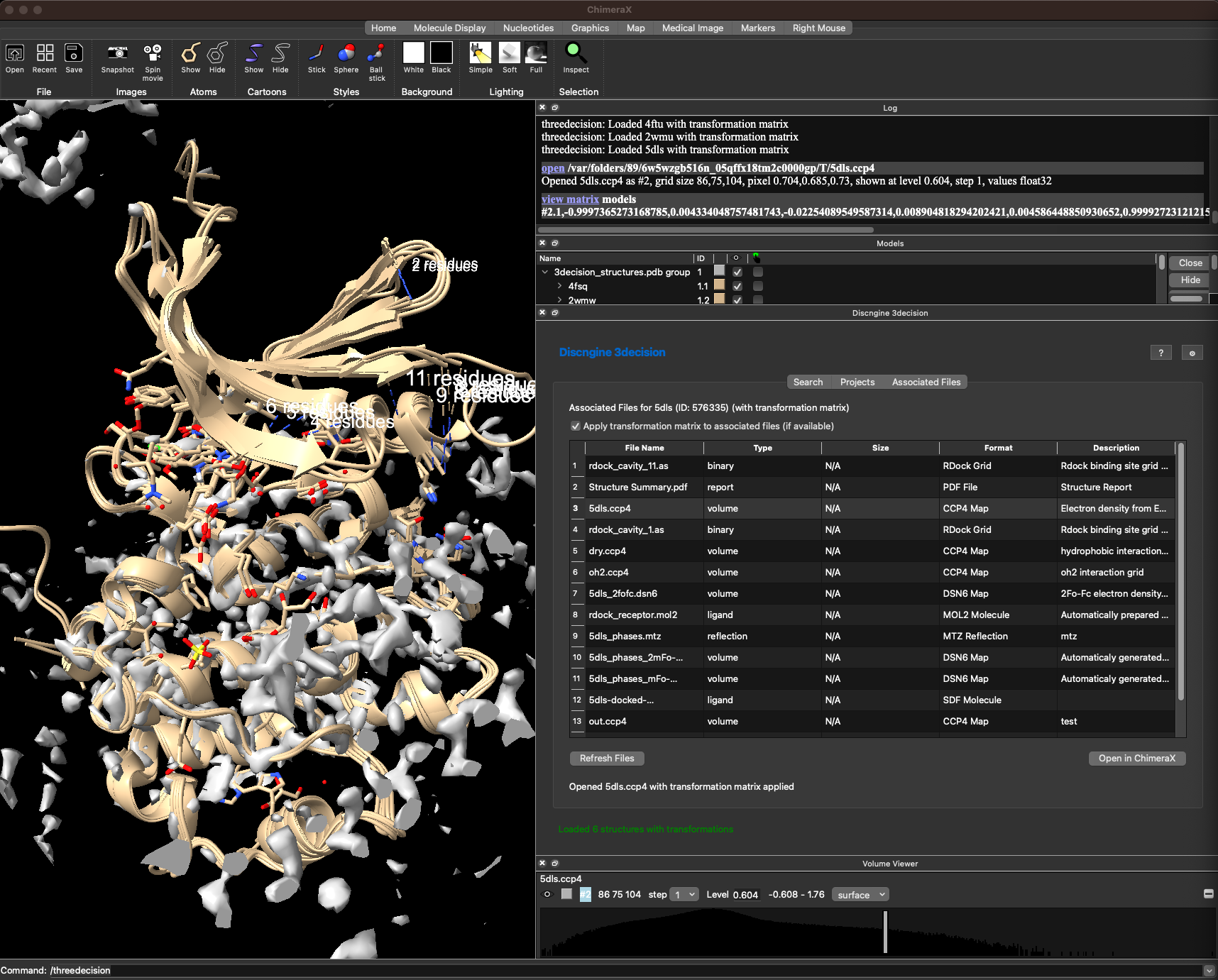
¶ License
This project is licensed under the MIT License.