¶ Introduction
3decision allows you to add both predicted and experimentally determined residue properties as sequence annotations. These data points (e.g. CDR regions, chemical liabilities, or alternative numbering schemes) are added as structure residue annotations and can be visualized directly in 3D on the resolved structure.
This tutorial walks you through the process of registering internal sequences, aligning them with structure chains, and adding sequence annotations to the correct residues. It will take you from the initial situation to the final outcome described below.
¶ Example Data
In this tutorial, we'll work on a protein structure of a Fab bound to an antigen.
To simulate an internal structure, we've registered this public protein structure using the 3decision user interface, with default settings. Instructions can be found here.
Once the structure is registered in 3decision, it will be given an external code. In our example it's hfzbcu, but the code of your structure will be different.
¶ Initial Situation
- In the information Browser, we can see that none of the structure chains have been mapped to a UniProt code or an in-house sequence identifier:
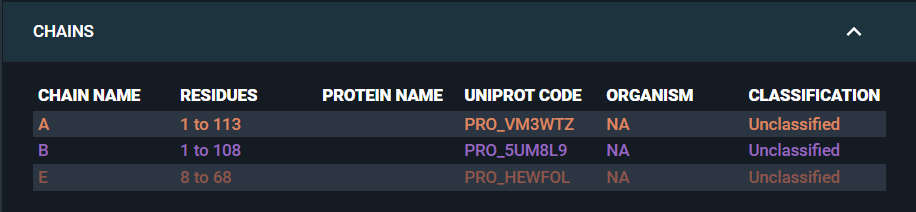
- In the Annotation Browser, we can see that no structure residue annotations have been added to the structure:
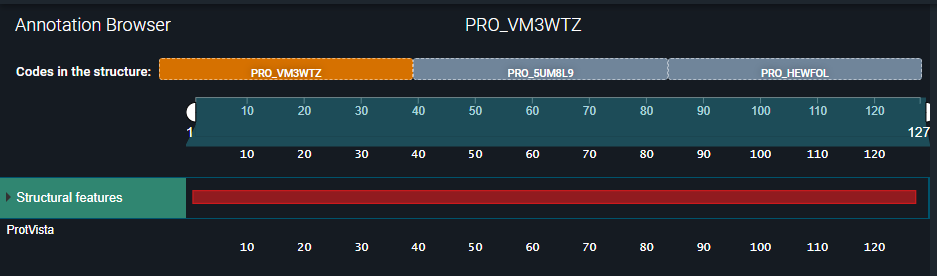

¶ Expected Outcome
- The antigen chain will be mapped to its corresponding UniProt identifier (CXL13_HUMAN) and the heavy and light Fab chains will be mapped to the corresponding heavy and light antibody chains:
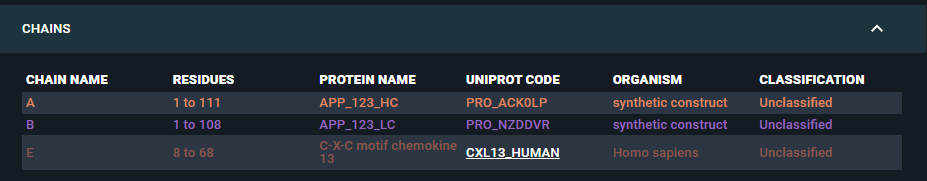
- The Fab CDR and predicted in-house residue properties will be registered as structure residue annotations:
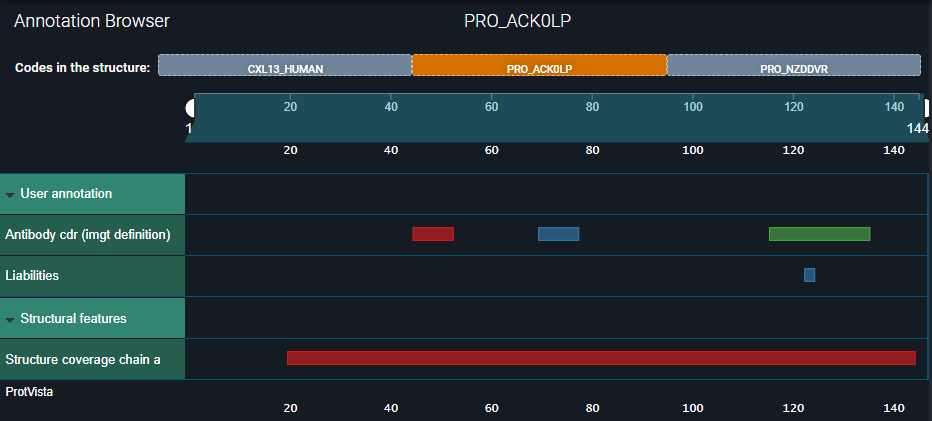
¶ Tutorial
¶ Step 1: Authenticate the User
To interact with the 3decision API endpoints, you must first set up API access by completing the following steps:
- Generate an API Secret Key
This key is required to request an access token and is generated manually via the 3decision UI. Instructions are available here. API Secret Keys may be configured to remain valid for a maximum duration of one year. - Generate an Access Token
Use the endpointGET /auth/api/loginto obtain a token that grants authorization. Access tokens are valid for up to one hour.
¶ Step 2: Register Internal Sequences
Register the full sequence of the Fab's heavy and light chains in 3decision using the API endpoint:
POST /sequence/register.
Assign each sequence a name that corresponds to its internal identifier, such as the "Actual Product Protein" identifier (e.g., APP_123_HC and APP_123_LC).
The taxonId 32630 corresponds to a synthetic construct. The parameters sequenceName and sequenceComment are optional.
More details on the usage of this endpoint can be found here.
Heavy Chain - curl command
curl -X 'POST' \
'https://3decision-[customer]-api.discngine.cloud/sequence/register' \
-H 'accept: application/json' \
-H 'X-API-VERSION: 1' \
-H 'Authorization: Bearer eyJhb********htE' \
-H 'Content-Type: application/json' \
-d '{
"proteinName": "APP_123_HC",
"geneName": "APP_123_HC",
"sequence": "MDWTWRFLFVVAAATGVQSQVQLVQSGAEVKKPGSSVKVSCKASGGTFSSYAISWVRQAPGQGLEWMGGIIPIFGTANYAQKFQGRVTITADESTSTAYMELSSLRSEDTAVYYCAREPDYYDSSGYYPIDAFDIWGQGTTVTV",
"sequenceName": "",
"sequenceVersion": 1,
"sequenceComment": "",
"taxonId": 32630,
"commit": true
}
Heavy Chain - API response body
{
"biomolecule": {
"proteinPrimaryName": "APP_123_HC",
"genePrimaryName": "APP_123_HC",
"organism": {
"name": "synthetic construct",
"id": 291123,
"taxonId": 32630,
"organismId": 291123,
"organismLabel": "synthetic construct",
"organismCode": 32630
},
"entryVersion": 1,
"biomoleculeId": 11209882,
"sourceId": 9003,
"biomoleculeCode": "PRO_ACK0LP",
"type": "Protein"
},
"biomoleculeSequenceId": 11286829,
"isCanonical": "1",
"sequenceVersion": 1,
"sequence": "MDWTWRFLFVVAAATGVQSQVQLVQSGAEVKKPGSSVKVSCKASGGTFSSYAISWVRQAPGQGLEWMGGIIPIFGTANYAQKFQGRVTITADESTSTAYMELSSLRSEDTAVYYCAREPDYYDSSGYYPIDAFDIWGQGTTVTV",
"sequenceLength": 144,
"biomoleculeSequenceName": "",
"biomolSequenceComment": ""
}
Light Chain - curl command
curl -X 'POST' \
'https://3decision-[customer]-api.discngine.cloud/sequence/register' \
-H 'accept: application/json' \
-H 'X-API-VERSION: 1' \
-H 'Authorization: Bearer eyJhb********htE' \
-H 'Content-Type: application/json' \
-d '{
"proteinName": "APP_123_LC",
"geneName": "APP_123_LC",
"sequence": "MAWALLLLTLLTQGTGSWAQSALTQPASVSASPGQSITISCTGTSSDVGAYDWVSWYQQHPGKAPKLLIFDVNNRPSGVSHRFSGSKSGNTASLTISGLQAEDEADYYCSSYTRRDTYVFGTGTKVTVLGENLYFQ",
"sequenceName": "",
"sequenceVersion": 1,
"sequenceComment": "",
"taxonId": 32630,
"commit": true
}
Light Chain - API response body
{
"biomolecule": {
"proteinPrimaryName": "APP_123_LC",
"genePrimaryName": "APP_123_LC",
"organism": {
"name": "synthetic construct",
"id": 291123,
"taxonId": 32630,
"organismId": 291123,
"organismLabel": "synthetic construct",
"organismCode": 32630
},
"entryVersion": 1,
"biomoleculeId": 11209883,
"sourceId": 9003,
"biomoleculeCode": "PRO_NZDDVR",
"type": "Protein"
},
"biomoleculeSequenceId": 11286830,
"isCanonical": "1",
"sequenceVersion": 1,
"sequence": "MAWALLLLTLLTQGTGSWAQSALTQPASVSASPGQSITISCTGTSSDVGAYDWVSWYQQHPGKAPKLLIFDVNNRPSGVSHRFSGSKSGNTASLTISGLQAEDEADYYCSSYTRRDTYVFGTGTKVTVLGENLYFQ",
"sequenceLength": 136,
"biomoleculeSequenceName": "",
"biomolSequenceComment": ""
}
The Fab heavy and light chains have now been registered under the biomoleculeCodes PRO_ACK0LP and PRO_NZDDVR, respectively.
¶ Step 3: Align Structure Chains with Sequences
When a biomoleculeCode — such as a UniProt code or a registered internal sequence — is provided during the structure registration process, the structure chains are aligned with the biomolecule's sequence. Each structure residue is then mapped to its corresponding sequence position. This analysis is referred to as Sequence mapping.
Before submitting our newly generated Fab biomoleculeCodes for sequence mapping, let's review the current mapping of our structure. This step helps us verify how many copies of the Fab and antigen are present, what their chain names are, and whether a UniProt code has already been assigned to the antigen.
¶ 3.a. Check Current Sequence Mapping
First, retrieve the internal ID of your structure using the API endpoint:
GET /structures/code/{external_code}.
Use the external_code you generated in the 'Example data' section of the Introduction (e.g. hfzbcu).
curl command
curl -X 'GET' \
'https://3decision-[customer]-api.discngine.cloud/structures/code/hfzbcu' \
-H 'accept: application/json' \
-H 'X-API-VERSION: 1' \
-H 'Authorization: Bearer eyJhb********htE'
API response body
{
"structures": [
{
"structure_id": 844553,
"external_code": "hfzbcu"
}
]
}
Next, inspect the current sequence mapping via the API endpoint GET /structures/info/chain providing the structure_id you retrieved in the previous step.
curl command
curl -X 'GET' \
'https://3decision-[customer]-api.discngine.cloud/structures/info/chain?structure_id=844553' \
-H 'accept: application/json' \
-H 'X-API-VERSION: 1' \
-H 'Authorization: Bearer eyJhb********htE'
API response body
{
"CHAINS_INFO": [
{
"BIOMOL_CODE_DISTINCT": [
"PRO_VM3WTZ",
"PRO_5UM8L9",
"PRO_HEWFOL"
],
"BIOMOL_CODE": [
"PRO_VM3WTZ",
"PRO_5UM8L9",
"PRO_HEWFOL"
],
"PROTEIN_PRIMARY_NAME_DISTINCT": [
null
],
"PROTEIN_PRIMARY_NAME": [
null,
null,
null
],
"GENE_PRIMARY_NAME_DISTINCT": [
null
],
"GENE_PRIMARY_NAME": [
null,
null,
null
],
"SPECIES_DISTINCT": [
"NA"
],
"SPECIES": [
"NA",
"NA",
"NA"
],
"SIZE_SORT_DISTINCT": [
127,
112,
61
],
"SIZE_SORT": [
127,
112,
61
],
"Chains": [
{
"CHAIN_CODE": "A",
"BIOMOL_ID": 11209876,
"BIOMOL_CODE": "PRO_VM3WTZ",
"PROTEIN_PRIMARY_NAME": null,
"GENE_PRIMARY_NAME": null,
"ORGANISM": "NA",
"Category": "Unclassified",
"ORGANISM_LABEL": "NA",
"STRUCTURE_ID": 844553,
"Range": "1-127",
"chainLength": 127
},
{
"CHAIN_CODE": "B",
"BIOMOL_ID": 11209877,
"BIOMOL_CODE": "PRO_5UM8L9",
"PROTEIN_PRIMARY_NAME": null,
"GENE_PRIMARY_NAME": null,
"ORGANISM": "NA",
"Category": "Unclassified",
"ORGANISM_LABEL": "NA",
"STRUCTURE_ID": 844553,
"Range": "128-239",
"chainLength": 112
},
{
"CHAIN_CODE": "E",
"BIOMOL_ID": 11209878,
"BIOMOL_CODE": "PRO_HEWFOL",
"PROTEIN_PRIMARY_NAME": null,
"GENE_PRIMARY_NAME": null,
"ORGANISM": "NA",
"Category": "Unclassified",
"ORGANISM_LABEL": "NA",
"STRUCTURE_ID": 844553,
"Range": "240-300",
"chainLength": 61
}
],
"STRUCTURE_ID": 844553
}
]
}
We can see in the response body that three chains - A, B and E - have been identified in the structure. They are currently mapped to three generic biomoleculeCodes: PRO_VM3WTZ, PRO_5UM8L9 and PRO_HEWFOL.
¶ 3.b. Submit New Sequence Mapping
Now, let's map each structure chain (A, B and E) to the appropriate biomoleculeCodes we generated in step 2 (PRO_ACK0LP and PRO_NZDDVR), along with the antigen's biomoleculeCode, which is the UniProt code CXL13_HUMAN.
This is done using the API endpoint POST /sequence-mapping with the payload shown below.
A successful call should return a 201 Created response from the server.
curl command
curl -X 'POST' \
'https://3decision-[customer]-api.discngine.cloud/sequence-mapping' \
-H 'accept: application/json' \
-H 'X-API-VERSION: 1' \
-H 'Authorization: Bearer eyJhb********htE' \
-H 'Content-Type: application/json' \
-d '{
"chains": [
{
"name": "A",
"biomolCode": "PRO_ACK0LP"
},
{
"name": "B",
"biomolCode": "PRO_NZDDVR"
},
{
"name": "E",
"biomolCode": "CXL13_HUMAN"
}
],
"externalCode": "hfzbcu",
"commit": true
}
Next, check the status of the sequence mapping analysis using the API endpoint:
GET /sequence-mapping/{externalCode}
curl command
curl -X 'GET' \
'https://3decision-[customer]-api.discngine.cloud/sequence-mapping/hfzbcu' \
-H 'accept: application/json' \
-H 'X-API-VERSION: 1' \
-H 'Authorization: Bearer eyJhb********htE'
API response body
{
"state": "success",
"content": {
"externalCode": "hfzbcu",
"createdBiomolStructure": 3,
"chains": [
{
"chainCode": "A",
"biomolAccession": "PRO_ACK0LP",
"createdSeqRes": 125,
"updateStrRes": 125
},
{
"chainCode": "B",
"biomolAccession": "PRO_NZDDVR",
"createdSeqRes": 112,
"updateStrRes": 112
},
{
"chainCode": "E",
"biomolAccession": "CXL13_HUMAN",
"createdSeqRes": 61,
"updateStrRes": 61
}
]
}
}
In the response body, we can see that the state of the analysis is success and that the three chains have been mapped to the submitted biomoleculeCodes.
¶ Step 4: Register Structure Residue Annotations
¶ 4.a. Retrieve Matching Structure Residue Positions
Now that your structure chains are correctly mapped to the original biomolecular sequences, we can easily verify to which sequence position (seq_res_number) each structure residue (str_res_number) corresponds. This step is essential if your custom annotations or properties were calculated on the original sequence rather than directly on the structure.
In this tutorial, we want to register four sequence annotations associated to the heavy chain sequence (registered in step 2):
- CDR1: residues 45–52
- CDR2: residues 70-77
- CDR3: residues 116-135
- Liability (Isomerization): residue 123-124
The numbering above refers seq_res_numbers. To retrieve the corresponding str_res_numbers endpoint str_res_numbers that we want to annotate, use the API endpoint:
GET /structures/structure/{external_code}/biomol/{biomol_code}/residues
Please refer the example below.
CDR2 residues - curl command
curl -X 'GET' \
'https://3decision-[customer]-api.discngine.cloud/structures/structure/hfzbcu/biomol/PRO_ACK0LP/residues?query_seq_res_numbers=70&query_seq_res_numbers=71&query_seq_res_numbers=72&query_seq_res_numbers=73&query_seq_res_numbers=74&query_seq_res_numbers=75&query_seq_res_numbers=76&query_seq_res_numbers=77' \
-H 'accept: application/json' \
-H 'X-API-VERSION: 1' \
-H 'Authorization: Bearer eyJhb********htE'
API response body
{
"seq_res_num": 70,
"external_code": "hfzbcu",
"biomolecule_code": "PRO_ACK0LP",
"str_res_num": "51",
"chain_code": "A"
},
{
"seq_res_num": 71,
"external_code": "hfzbcu",
"biomolecule_code": "PRO_ACK0LP",
"str_res_num": "52",
"chain_code": "A"
},
{
"seq_res_num": 72,
"external_code": "hfzbcu",
"biomolecule_code": "PRO_ACK0LP",
"str_res_num": "52A",
"chain_code": "A"
},
{
"seq_res_num": 73,
"external_code": "hfzbcu",
"biomolecule_code": "PRO_ACK0LP",
"str_res_num": "53",
"chain_code": "A"
},
{
"seq_res_num": 74,
"external_code": "hfzbcu",
"biomolecule_code": "PRO_ACK0LP",
"str_res_num": "54",
"chain_code": "A"
},
{
"seq_res_num": 75,
"external_code": "hfzbcu",
"biomolecule_code": "PRO_ACK0LP",
"str_res_num": "55",
"chain_code": "A"
},
{
"seq_res_num": 76,
"external_code": "hfzbcu",
"biomolecule_code": "PRO_ACK0LP",
"str_res_num": "56",
"chain_code": "A"
},
{
"seq_res_num": 77,
"external_code": "hfzbcu",
"biomolecule_code": "PRO_ACK0LP",
"str_res_num": "57",
"chain_code": "A"
}
After verifying all positions, we find that the corresponding str_res_numbers are as follows:
- CDR1: residues 26, 27, 28, 29, 30, 31, 32, 33
- CDR2: residues 51, 52, 52A, 53, 54, 55, 56, 57
- CDR3: residues 93, 94, 95, 96, 97, 98, 99, 100, 100A, 100B, 100C, 100D, 100E, 100F, 100G, 100H, 100I, 100J, 101, 102
- Liability (Isomerization): residues 100, 100A
¶ 4.b. Register Annotation Labels
Before registering the structure residue annotations, we need to register the annotation label under which the annotations will be grouped visually in the Annotation Browser of the user interface. Annotation labels are registered via the API endpoint:
POST /annotation-types.
CDRs - curl command
curl -X 'POST' \
'https://3decision-[customer]-api.discngine.cloud/annotation-types' \
-H 'accept: application/json' \
-H 'X-API-VERSION: 1' \
-H 'Authorization: Bearer eyJhb********htE' \
-H 'Content-Type: application/json' \
-d '{
"annot_type_label": "Antibody CDR (IMGT definition)"
}
Liabilities - curl command
curl -X 'POST' \
'https://3decision-[customer]-api.discngine.cloud/annotation-types' \
-H 'accept: application/json' \
-H 'X-API-VERSION: 1' \
-H 'Authorization: Bearer eyJhb********htE' \
-H 'Content-Type: application/json' \
-d '{
"annot_type_label": "Liabilities"
}
If the annotation label is successfully registered, the API call will return a 201 Created response.
If the annotation label already exists in the database, the call will return a 400 Bad Request response.
You can retrieve a list of all registered annotation labels using the API endpoint:
GET /annotation-types.
¶ 4.c. Register the Annotations
Everything is now ready to register the structure residue annotations using the API endpoint:
POST /annotations/structure-residues.
A full description of this step is available in the documentation here.
curl command
curl -X 'POST' \
'https://3decision-[customer]-api.discngine.cloud/annotations/structure-residues' \
-H 'accept: application/json' \
-H 'X-API-VERSION: 1' \
-H 'Authorization: Bearer eyJhb********htE' \
-H 'Content-Type: application/json' \
-d '{
"annotations": [
{
"annotation": {
"label": "CDR-H1",
"typeLabel": "Antibody CDR (IMGT definition)",
"source": "MOE",
"link": "",
"groups": null,
"data": null
},
"location": [
{
"authResNum": 26,
"authChain": "A",
"insertionCode": ""
},
{
"authResNum": 27,
"authChain": "A",
"insertionCode": ""
},
{
"authResNum": 28,
"authChain": "A",
"insertionCode": ""
},
{
"authResNum": 29,
"authChain": "A",
"insertionCode": ""
},
{
"authResNum": 30,
"authChain": "A",
"insertionCode": ""
},
{
"authResNum": 31,
"authChain": "A",
"insertionCode": ""
},
{
"authResNum": 32,
"authChain": "A",
"insertionCode": ""
},
{
"authResNum": 33,
"authChain": "A",
"insertionCode": ""
}
]
},
{
"annotation": {
"label": "CDR-H2",
"typeLabel": "Antibody CDR (IMGT definition)",
"source": "MOE",
"link": "",
"groups": null,
"data": null
},
"location": [
{
"authResNum": 51,
"authChain": "A",
"insertionCode": ""
},
{
"authResNum": 52,
"authChain": "A",
"insertionCode": ""
},
{
"authResNum": 52,
"authChain": "A",
"insertionCode": "A"
},
{
"authResNum": 53,
"authChain": "A",
"insertionCode": ""
},
{
"authResNum": 54,
"authChain": "A",
"insertionCode": ""
},
{
"authResNum": 55,
"authChain": "A",
"insertionCode": ""
},
{
"authResNum": 56,
"authChain": "A",
"insertionCode": ""
},
{
"authResNum": 57,
"authChain": "A",
"insertionCode": ""
}
]
},
{
"annotation": {
"label": "CDR-H3",
"typeLabel": "Antibody CDR (IMGT definition)",
"source": "MOE",
"link": "",
"groups": null,
"data": null
},
"location": [
{
"authResNum": 93,
"authChain": "A",
"insertionCode": ""
},
{
"authResNum": 94,
"authChain": "A",
"insertionCode": ""
},
{
"authResNum": 95,
"authChain": "A",
"insertionCode": ""
},
{
"authResNum": 96,
"authChain": "A",
"insertionCode": ""
},
{
"authResNum": 97,
"authChain": "A",
"insertionCode": ""
},
{
"authResNum": 98,
"authChain": "A",
"insertionCode": ""
},
{
"authResNum": 99,
"authChain": "A",
"insertionCode": ""
},
{
"authResNum": 100,
"authChain": "A",
"insertionCode": ""
},
{
"authResNum": 100,
"authChain": "A",
"insertionCode": "A"
},
{
"authResNum": 100,
"authChain": "A",
"insertionCode": "B"
},
{
"authResNum": 100,
"authChain": "A",
"insertionCode": "C"
},
{
"authResNum": 100,
"authChain": "A",
"insertionCode": "D"
},
{
"authResNum": 100,
"authChain": "A",
"insertionCode": "E"
},
{
"authResNum": 100,
"authChain": "A",
"insertionCode": "F"
},
{
"authResNum": 100,
"authChain": "A",
"insertionCode": "G"
},
{
"authResNum": 100,
"authChain": "A",
"insertionCode": "I"
},
{
"authResNum": 100,
"authChain": "A",
"insertionCode": "J"
},
{
"authResNum": 101,
"authChain": "A",
"insertionCode": ""
},
{
"authResNum": 102,
"authChain": "A",
"insertionCode": ""
}
]
},
{
"annotation": {
"label": "Isomerization",
"typeLabel": "Liabilities",
"source": "NaturalAntibody",
"link": ""
},
"location": [
{
"authResNum": 100,
"authChain": "A",
"insertionCode": ""
},
{
"authResNum": 100,
"authChain": "A",
"insertionCode": "A"
}
]
}
],
"externalCode": "hlt3av",
"commit": true
}
If the request is successful, the response will include an annotation ID for each annotation that was registered:
API response body
[
7458482,
7458483,
7458484,
7458485
]
¶ Final Result
¶ Retrive Results using the API
All registered sequence annotations for a given biomolecule–structure pair can be retrieved using the following API endpoint:
GET /annotations/annotation/{biomol_code}/{external_code}
This endpoint also returns the full biomolecule sequence, along with an additional annotation that describes which portion of the sequence is resolved in the structure.
This type of annotation has the CATEGORY value "STRUCTURAL".
curl command
curl -X 'GET' \
'https://3decision-[customer]-api.discngine.cloud/annotations/annotation/PRO_ACK0LP/hfzbcu' \
-H 'accept: application/json' \
-H 'X-API-VERSION: 1' \
-H 'Authorization: Bearer eyJhb********htE'
}
API response body
{
"sequence": "MDWTWRFLFVVAAATGVQSQVQLVQSGAEVKKPGSSVKVSCKASGGTFSSYAISWVRQAPGQGLEWMGGIIPIFGTANYAQKFQGRVTITADESTSTAYMELSSLRSEDTAVYYCAREPDYYDSSGYYPIDAFDIWGQGTTVTV",
"features": [
{
"ftid": "ANNOT_ID_7458482",
"begin": 45,
"end": 52,
"description": "CDR-H1",
"type": "Antibody CDR (IMGT definition)",
"source": "MOE",
"created_by": "gabriella.jonasson@discngine.com",
"created_date": "2025-07-31T15:27:18.000Z",
"category": "user_annotation",
"color": "#e41a1c"
},
{
"ftid": "ANNOT_ID_7458483",
"begin": 70,
"end": 77,
"description": "CDR-H2",
"type": "Antibody CDR (IMGT definition)",
"source": "MOE",
"created_by": "gabriella.jonasson@discngine.com",
"created_date": "2025-07-31T15:27:18.000Z",
"category": "user_annotation",
"color": "#377eb8"
},
{
"ftid": "ANNOT_ID_7458484",
"begin": 116,
"end": 135,
"description": "CDR-H3",
"type": "Antibody CDR (IMGT definition)",
"source": "MOE",
"created_by": "gabriella.jonasson@discngine.com",
"created_date": "2025-07-31T15:27:18.000Z",
"category": "user_annotation",
"color": "#4daf4a"
},
{
"ftid": "ANNOT_ID_7458485",
"begin": 123,
"end": 124,
"description": "Isomerization",
"type": "Liabilities",
"source": "NaturalAntibody",
"created_by": "gabriella.jonasson@discngine.com",
"created_date": "2025-07-31T15:27:18.000Z",
"category": "user_annotation",
"color": "#377eb8"
},
{
"ftid": "ANNOT_ID_CHAIN_A_0",
"begin": 20,
"end": 144,
"description": "chain A resolved on structure",
"type": "structure_coverage_chain_A",
"source": null,
"created_by": null,
"created_date": null,
"category": "STRUCTURAL",
"color": "#e41a1c"
}
]
}
¶ Visualize Results in the User Interface
¶ Sequence Mapping
Under the Information Browser, you can now see the updated sequence mapping:
- The Fab's heavy (A) and light (B) chains are mapped to the corresponding antibody biomolecule sequences, each labeled with the name of its internal sequence database reference: APP_123_HC and APP_123_LC.
- The antigen chain (E) is mapped to its corresponding UniProt identifier: CXL13_HUMAN.

¶ Sequence annotations
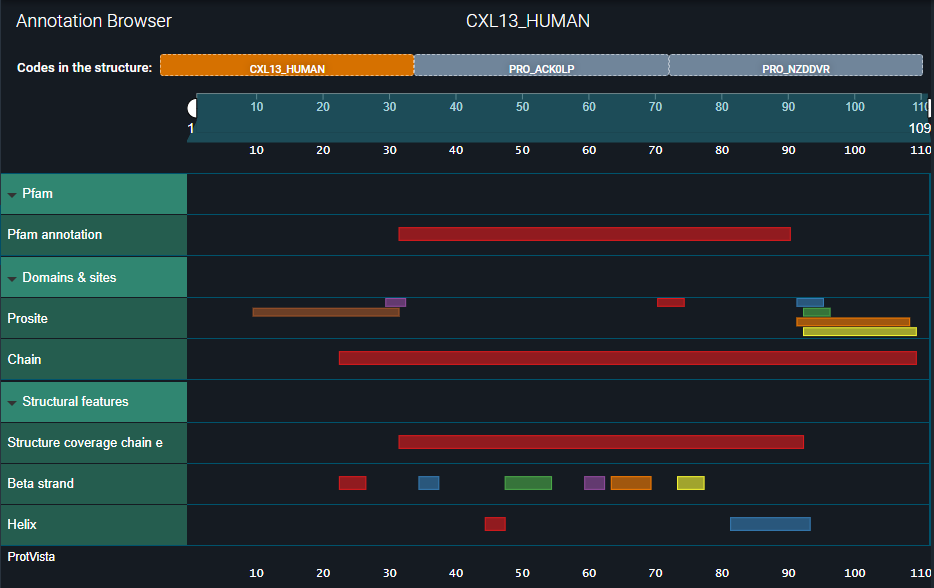
Under the Annotation Browser, you can now see the three updated biomolecule codes to which the structure chains are mapped:
- In the first one — CXL13_HUMAN — you can see all annotations inherited from the UniProt reference.
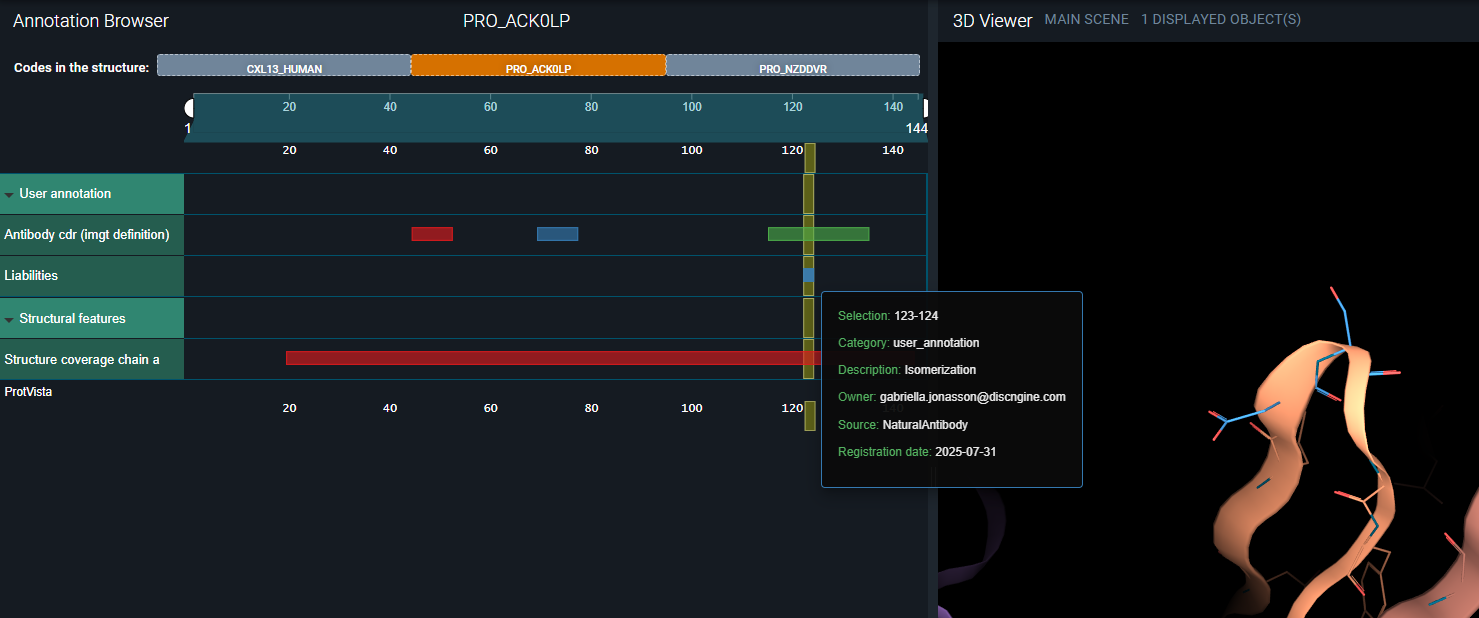
- In the second one — PRO_ACK0LP — you can see the annotations we registered for the heavy chain.
- In the third one — PRO_NZDDVR — only the structural coverage of the light chain is visible.
See the three images below.
¶

¶
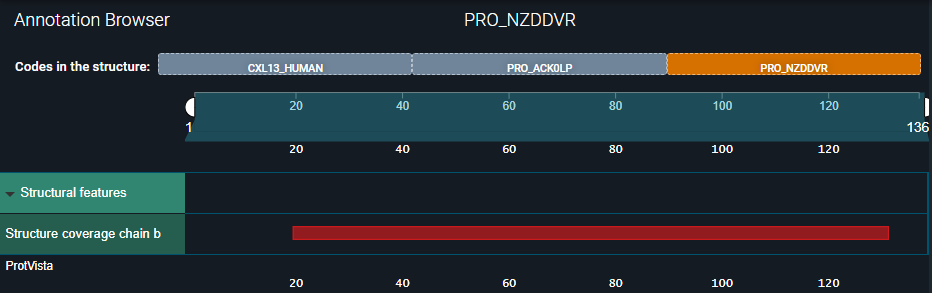

Details about the registered sequence annotations are displayed when you hover over an annotation in the Annotation Browser: